National and international collaborations are important for NCMM, as we seek to strengthen our connections with Norway’s university and research environments outside of Oslo and beyond.
To highlight some of these connections, we caught up with Marieke Kuijjer, head of the Computational Biology and Systems Medicine Group at NCMM, to hear about her work with NCMM Board Member, Professor Ola Myklebost, based at the University of Bergen, and her links to Leiden University, Netherlands
You have previously collaborated with NCMM’s Board member, Ola Myklebost, a Professor at the University of Bergen on some projects. Can you tell me a bit more about how you came to work together?
I have known Ola for some time – we collaborated during my PhD as we were part of the same European network of excellence, EuroBoNet. In terms of our more recent work together, Ola led a project that centred around a large cancer drug screen. This involved testing drugs, either that were FDA approved or that were already in clinical trials, against liposarcoma - a very rare cancer type. This work was based on 13 cell lines; quite a large amount for such a rare cancer. Ola’s group conducted the drug screen and found a couple of drugs that they wanted to further test. They did some experiments to see how well these drugs could kill cancer cells and what the mechanisms were behind this. Since then they have tested two promising drugs in vivo, in patient-derived xenografts in mice, and one of them has also proved very efficient there.
The initial research by Ola’s group found that the response across these 13 cell lines was quite variable. This led to the question of whether it would be possible to find biomarkers to indicate what these cell lines’ responses to the cancer drugs might be. If the cell lines’ responses to these tested drugs were quite variable, then the response would likely be variable in patients too. One of Ola’s postdocs has bioinformatics expertise and had conducted a bioinformatics analysis to look into the expected targets of these drugs. She found that they didn’t correlate well with drug responses across the cell lines. This is when Ola approached me to ask if I could join the project and conduct a genome-wide screen to help with the biomarker analysis.
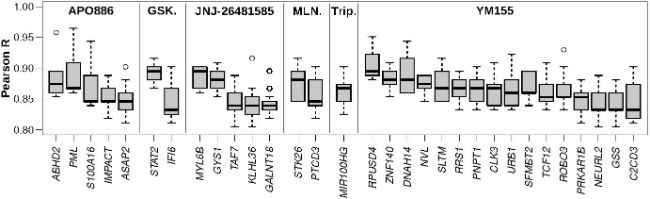
We screened all genes and then checked if they were associated with drug response. Although the known drug targets did not correlate particularly well, we found that with each of the drugs there were a handful of genes that did correlate favourably and robustly. The pathways that these responses were working in also made sense. For example, there was one gene that played a role in glycolysis that was a biomarker for a response to histone deacetylase inhibitors. This was something that we expected, as we already knew that the pathway here was known to play a role in histone acetylation.
Are there any next steps to further explore these biomarkers?
In terms of where things go from here, we hope that these findings will one day be tested in the clinic. The drugs used in the screen are all approved for cancer but not yet for sarcomas. Some have quite nasty side effects and so need to be fully clinically tested to be deemed safe and effective for sarcoma patients. We hope that the biomarkers we identified can be tested in a clinical setting and so used eventually to see which patients might respond to a particular drug.
Are there other sarcoma projects that you and Ola have worked on in the past?
Ola initiated NoSarC (Norwegian Sarcoma Consortium), which is a collaborative research project. The project has enabled sarcoma scientists and clinics across Norway to develop a bio and databank of different sarcomas gathered from across the country. With the genome and clinical data available through NoSarC, we may conduct further analysis using a tool developed by my postdoc Tatiana (Belova). The tool describes heterogeneity in a patient population in terms of gene regulation. Tatiana has applied the tool to different types of leiomyosarcoma and found some interesting pathways. We are now at the stage where we’re ready to validate some of our findings. The data collected by Ola as part of NoSarC is therefore perfect for this. It’s certainly different from the work I mentioned previously with the cell lines as it doesn’t involve drug responses, however, many of the techniques our group uses are similar.
How will you work to further explore your initial findings?
In terms of what comes next, a logical step to extend this work would be to experimentally validate the heterogeneity of gene regulation in these pathways. Tatiana has already validated some of these pathways in an independent leiomyosarcoma cohort. This has been in collaboration with Priya Chudasama at the German Cancer Research Center (DKFZ), together with whom we will also perform functional validation using leiomyosarcoma cell lines.
You recently became an adjunct professor at Leiden University. Can you tell me a bit more about this and what the plans for the future are in terms of your association?
I’ve had a long-standing connection with Leiden, having done my PhD there myself, and having supervised a visiting PhD student from Leiden during my time in Boston. After being invited to be an opponent in this student’s PhD defence, I gave a talk about my research at NCMM. I learnt after this that the University wanted to strengthen its expertise in computational oncology research and they realized that I could positively contribute towards this. This then grew into a more formal association for me. We will work on profiling multi-omic data types, including gene regulatory networks, and look into many different types of sarcoma. This links back nicely to the work I have already done with Ola (Myklebost).
The plan is to look into and profile the sarcoma once it has been diagnosed and then also when it recurs or metastasizes. We will examine how the tumour develops and then we will measure different data types with whole-genome sequencing and RNA sequencing. We will also model gene regulatory networks and see how they evolve through tumour development. Together, these data types will contribute to forming ‘digital twins’ for individual patients and our network analysis will be part of this. I’m also now co-supervising a PhD student at Leiden and, hopefully, once travel becomes easier again, we will be able to organize more visits between us. One positive thing about COVID and the restrictions is that the PhD student has been able to join our group meetings and we’ve all felt a bit more connected this way.
Your group has grown over the past 12 months. Can you describe some of the projects they have been working on and how things are progressing for them?
We’ve had a busy time despite COVID restrictions and the group has been doing well. We recently published two review papers, one written by Genís, (Calderer i García) a Research Assistant in the group, which concerns network module detection approaches. This was published in Frontiers in Genetics (https://doi.org/10.3389/fgene.2021.649440). Debora (Meijer), the PhD student from Leiden University, wrote a review on immunogenomics in sarcoma, which has been published in the journal Biomedicines (https://doi.org/10.3390/biomedicines9081048).
We have a recent BioRxiv pre-print that describes a tool developed by our PhD student, Ping-Han (Hsieh). Ping-Han developed a tool that normalizes data ahead of modelling a network. It’s currently up at: Adjustment of spurious correlations in co-expression measurements from RNA-Sequencing data. Daniel (Osorio), postdoc in the group, developed an R package that helps with downloading and merging single-cell data which has been published in the journal Bioinformatics (https://pubmed.ncbi.nlm.nih.gov/34320637/). Daniel has also developed a computational approach to recommend treatment combinations for cancer through single-cell transcriptomic data analysis and is currently drafting a manuscript to describe the method together with an analysis in breast cancer.
Tatiana’s previously-mentioned tool that looks into patient heterogeneity of gene regulation is currently being written up into a manuscript and will shortly be submitted. We also have a network analysis that has now been accepted into the journal Cancer Research. This describes analysis in glioblastoma (brain cancer) where we found that the regulation of PD1 signalling is associated with glioblastoma survival and tumour aggressiveness.

expression/targeting in short-term survivors, blue: higher expression/targeting in the long-term survivors. Image: Kuijjer/Pop
We’re also working hard to get our ‘Pink Ribbon’ breast cancer project, funded by Kretforeningen, started. Our PhD student, Romana (Pop), has already started on some of the method development for the project. She’s trying to integrate different types of ‘omics data with the gene regulatory network models that we have. We will also hire a PhD candidate for the data analysis part of the project who can help to progress things here. We’ll share more news and updates with regards to these projects soon – watch this space.
Learn more about the Kuijjer group at their NCMM web page: https://www.med.uio.no/ncmm/english/groups/kuijjer-group/
And at their Kuijjer lab page at: https://www.kuijjerlab.org