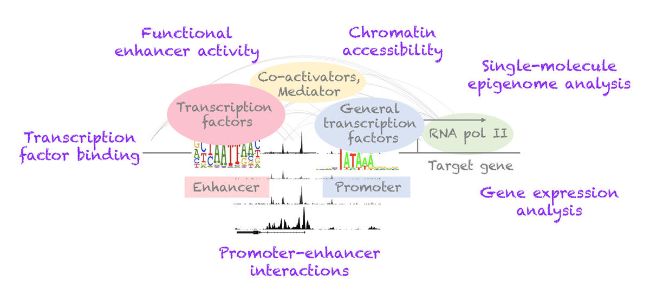
We develop innovative experimental methods for studying gene regulation, enhancer function and epigenome reprogramming, including cell fate conversion methods and functional genomics at single-cell and single-molecule resolution.
Research focus
Recurrent mutations have been catalogued in all major human tumour and cancer types. It is known that the same mutations do not cause cancer in all tissues and that mutations also accumulate in normal tissues not developing cancer. However, it is still poorly understood, why a certain tissue is vulnerable to distinct oncogenic mutations, but others are not.
Our goal is to understand the molecular mechanisms that lead to organ-specific cancer with a special focus on the role of lineage-specific transcription factors in early tumorigenic events. We utilize a systems biology approach with a plethora of cutting-edge experimental methods to study enhancer malfunction, genome plasticity and cancer-specific gene regulation aiming for precision targeting of malfunctioning gene regulatory elements in human cancer.
Main projects
1. Molecularly controlled genomics approach for studying early tumorigenic events
We have developed a novel cell fate conversion approach for elucidating the molecular events leading to pancreatic cancer. The current focus is on studying the precise determinants of pancreatic cell identity and oncogenic transformation at single-cell resolution using multi-layered omics data for chromatin state, transcription factor binding and gene expression programs.
2. Enhancer malfunction and epigenetic reprogramming in lineage-specific cancer
Transcription factors control tissue- and context-specific gene expression through regulatory elements such as promoters and enhancers. Our innovative multicancer–multiomics approach aims to identify precise enhancer reprogramming events that cancer cells hijack for activating tumorigenic signalling pathways. For this, we are using a wide range of experimental methods from massively parallel reporter assays to genome-wide analysis of transcription factor binding and from CRISPR-based genome editing to single molecule epigenome analysis using long-read nanopore sequencing. Using this wide repertoire of modern state-of-the-art next-generation methods, the specific areas of focus in the laboratory are:
- Nuclear receptors and pioneer factors in epigenome reprogramming of hormonal cancers and disease progression
- Epigenetic plasticity and lineage-specificity of endodermal origin cancers
- Role of non-coding regulatory elements such as the transposable elements in driving oncogenic signalling in human cancers.
Impact
Many human cancers still lack specific diagnostic markers and effective treatment options. Pancreatic cancer, for example, is one of the most lethal cancer types that is typically diagnosed at late stages. The goal of our research is to identify specific regulatory features that promote tumorigenesis of endodermal-origin tissues, such as pancreatic cancer. Better understanding of the early tumorigenic events can lead to finding of novel biomarkers or tissue-specific vulnerabilities for developing targeted therapies.
Learn more
Visit the Sahu group’s external website for more information