Some of the diseases are caused by yet unknown infectious agents, clonal genetic mutations acquired later in life, or other environmental factors. Others are caused by hereditary single gene defects in pathways important for immune cell function.
In hereditary immune diseases, a complex interplay between environment and other genetic variants exist, and leads to a broad array of clinical diseases. The penetrance is often incomplete, meaning that one family member can get a very severe disease, whereas another stays asymptomatic. The same mutation can also lead to seemingly opposed phenotypes. One patient can develop a life-threatening, multisystemic autoimmune disease, whereas another with the same mutation might suffer from recurrent infections.
Optimisation of CRISPR-Cas gene editing to treat and diagnose rare immunological diseases
We aim to optimize the CRISPR-Cas gene editing technology, such as gene therapy in these diseases, and stratify the existing therapies. Current therapies include biologic drugs and small molecule inhibitors which aim to better target the underlying molecular pathways of disease.
Our group is interested in rare and undiagnosed immunological diseases that mainly present as autoimmunity, severe allergy, infection susceptibility, and early cancer. Some of the diseases are acquired and resemble malignant hematological diseases, whereas others are caused by germline single gene defects in pathways important for immune cell function. Even in genetic diseases, a complex interplay between environment and other genetic variants exists and leads to a broad array of clinical diseases. As an example, one patient can present with a severe blood vessel inflammation, whereas another with the same mutation might instead suffer from recurrent infections.
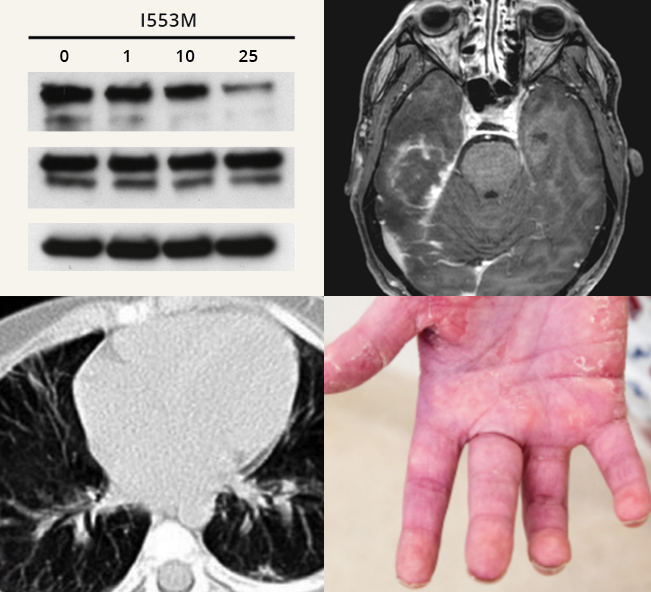
Genome editing for monogenic immune diseases
CRISPR-Cas9 is a new gene editing technology that can cut the genome in designated locations. In patients with genetic diseases, CRISPR-Cas9 can cut the disease-causing mutation, followed by repairing of the cells at the cut site to a normal sequence, provided that the cells have access to a repair DNA. While the cutting is efficient, the following repair is inefficient, and usually silences the gene.
Our goal is to improve the precision mutation correction by altering the pathways that govern DNA repair. We have attached DNA repair proteins to Cas9 to bias editing and carry the correct repair DNA sequence, and timed editing with the cell cycle. Our long-term goal is to develop a platform that can take any patient mutation and correct it to a “healthy” sequence, thus treating a large number of genetic diseases. Besides gene therapy, the platform could be adapted for other applications such as cell editing for cancer immunotherapy.
Personalized use of targeted therapies in rare diseases
Many genetic immune diseases harbor inherited mutations in pathways that can be targeted with novel therapies originally developed for cancer and autoimmune diseases. As an example, JAK inhibitor Ruxolitinib can target activating STAT3 mutations that cause early-onset multi-organ autoimmune disease. Compared to cell therapies and gene editing, the use of targeted drugs is a much faster and safer way to improve patient symptoms and quality of life.
For mutant genes that localize to known immunological pathways, the choice of targeted drug is often easy. However, in several mutations, the function of the gene is poorly known. One of our future projects is to systematically catalogue poorly characterized mutant protein interactomes in immune cells, and use the information to rationally choose drugs for diseases in which no obvious druggable targets exist.
In many patients with suspected genetic immunological disease, the mutation that causes their disease is either undruggable or not found despite extensive genome-wide sequencing studies. Our second future goal is to utilize transcriptome profiling to detect immunological profiles that the new therapies can target, and tailor the treatment based on the sequencing result. This would enable the rational use of biologics and small molecule inhibitors in patients who either lack a diagnosis or have a genetic mutation that can’t be directly targeted with existing therapies.
